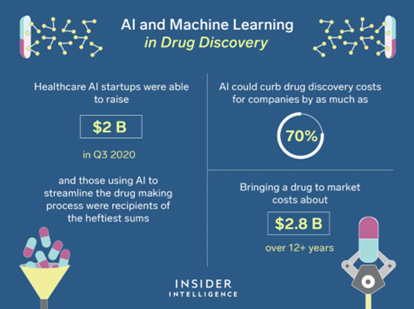
Finding a new treatment is more expensive and takes longer than at any other time in history. In recent years, the typical cost of bringing a new medication to market has climbed to almost $2.5 billion. The industry spent an additional $5 billion each year on R&D, while the number of approved drugs per year fell from 50 (1996) to approximately 15 (2010). For every molecule which becomes a medicine, millions from the same batch could be physically evaluated and eliminated as unfit.
A natural solution to the high failure rate of designed-drug molecules would be to make more informed choices about which ones to develop and test. How can we design molecules specifically for the task? More importantly, can we program an intelligent computing system to design it for us? These two ideas form the core philosophy of Computer Aided Drug Design (CADD).
To begin the CADD process, we need to identify the protein that is known to be related to a disease that we want to cure. This protein is referred to as a “target” in the life sciences. Proteins that promote inflammation, proteins that aid tumour growth and proteins that viruses utilise to infect human cells are all examples of therapeutic targets. The objective of pharmacological development is to develop chemical compounds that strongly interact with these targets, lowering (or boosting) their impact. These compounds are referred to as “ligands” for that target.
The 3D structure of the protein targets could interact with ligands of suitable shape, just like the correct key shape can fit and open a specific lock. Naturally, just like a professional lock-picker would study the structure of a lock to produce a duplicate key, we would also need to look at the 3D structure of a protein target to create a synthetic chemical ligand. In essence, we will look to the composition of the target protein to guide our predictions, and this is called the structure-based method.

Thus, it is important to have an algorithm that can perform a database search and suggest ligands that can bind a particular protein, inhibit enzyme activity, bind nucleic acid targets and finally predict binding modes of molecule-protein complexes. Next in the pipeline, the identified ligands will be fed into a deep learning classifier that can find the similarity of this molecule to the structure space covered by natural products. This will point out the suitable natural products that can achieve a good fit with the protein target. Protein folding and docking simulations via AlphaFold and AutoDock will be done at a later stage for validation.