Graphs may be found everywhere around us. Your social network is a network of individuals and connections. Think about it, at home you may be a parent with a spouse and a couple of children. At the same time, you are the middle child of 5 siblings. Additionally, you are an employee in a company with 300 personnel. At the weekends, you play football with your team, and you regularly hang out in a pub with a close circle of friends. Thus, if we plot your human relations with all individuals, your graph will look quite complex.
A graph may be used to represent anything that is made up of linked items. Graphs are great for visualising relationships between people, things, and concepts. Graphs can also be used to train machine learning models for complex tasks in addition to displaying information.
GNNs are a type of machine learning algorithm that can extract crucial information from graphs and generate meaningful predictions. GNNs have become a strong tool for many critical applications as graphs have gotten more prevalent and information-rich, and artificial neural networks have become more popular and capable.
Technically speaking, a graph is made up of nodes and edges. In a social network, for example, nodes can represent people and their features (e.g., name, gender, age, city), whereas edges might represent the users’ relationships. Other sorts of nodes, such as cities, sports teams, and news outlets, can be included in a more complicated social graph, as well as edges that define the relationships between the users and those nodes.
For our company, we deal with the graph that connects natural plants (together with their active chemical compounds) with target proteins (and related human illnesses). As one could imagine, each medicinal plants probably have multiple active chemical compounds that can be extracted from different parts of the plant. Similarly, a target protein could play important roles in multiple human maladies. Some of the active chemical compounds from different plant extracts could also cause an adverse reaction in a patient. Thus, to capture complex information for all natural medicinal plants, a graph structure would be more suitable instead of a tabular form.
Take for example, we start building a simple graph of disease. So, each different type of disease will occupy each unique node. Inside each node, we have necessary information related to its unique diseases such as typical target proteins, therapeutic chemical compounds, and other info. Some diseases may be connected, so they would form edges between their nodes. The information contained in each node can be summarised via a table. On the other hand, the graph’s overall connectedness may be represented as an adjacency matrix, which reveals which nodes are related to one another.

So how can we train a neural network to read this complicated graph structure? The GNN takes as input structured graph data and outputs a vector of numerical values representing important information about nodes and their relationships.
This type of vector representation is known as “graph embedding.” In machine learning, embeddings are frequently employed to turn complex information into a structure that can be discriminated against and learnt. Natural language processing systems, for example, employ word embeddings to generate numerical representations of words and their relationships.
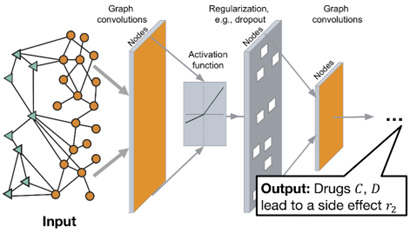
So, what are the benefits of GNN after it has learned the complex graph data? Well, the most important is to discover the potential connection of nodes via their shared attributes within the nodes. For example, the GNN might point out that two different types of diseases (at two different nodes) are very closely related via their shared target proteins. This is called clustering and it is the focus of most of our analysis. This will open up the possibility of repurposing previously tested chemical compounds for new diseases.